About
Biography
I’m a researcher interested in statistical genetics, complex traits, statistical rigor and allergic disorders. I work with various collaborators to evaluate how genetics may contribute to patient outcomes. These outcomes could be one or more allergic conditions or how an individual recovers after an injury.
I have always loved numbers, and the field of genetics fascinates me. Working in the field of statistical genetics allows me to combine these interests. I have been a researcher for over 25 years and began working at Cincinnati Children’s in 2002.
I am very proud of my work on congenital heart malformations. We have shown that these malformations can be due to the combinations of multiple genes supporting the role of complex inheritance. Beyond genetics, I’m excited about the opportunity to ensure that we reach out to other communities to share our work.
BA: University of Kansas, Lawrence, KS, 1991
MA: University of Kansas, Lawrence, KS, 1993
PhD: University of Kansas, Lawrence, KS, 1999
Fellowship: Post-doctoral, Southwest Foundation for Biomedical Research, San Antonio, TX, 2002
Services and Specialties
Interests
Statistical genetics; genetics of heart malformations; pharmacogenetics; genetics of allergic disorders
Research Areas
Publications
One-food versus six-food elimination diet therapy for the treatment of eosinophilic oesophagitis: a multicentre, randomised, open-label trial. The Lancet Gastroenterology and Hepatology. 2023; 8(5):408-421.
International Consensus Recommendations for Eosinophilic Gastrointestinal Disease Nomenclature. Clinical Gastroenterology and Hepatology. 2022; 20(11):2474-2484.e3.
Multiancestral polygenic risk score for pediatric asthma. Journal of Allergy and Clinical Immunology. 2022; 150(5):1086-1096.
The genetic architecture of pediatric cardiomyopathy. American Journal of Human Genetics. 2022; 109(2):282-298.
Novel role for caspase recruitment domain family member 14 and its genetic variant rs11652075 in skin filaggrin homeostasis. Journal of Allergy and Clinical Immunology. 2022; 149(2):708-717.
Desmoplakin and periplakin genetically and functionally contribute to eosinophilic esophagitis. Nature Communications. 2021; 12(1):6795.
Sensitization to peanut, egg or pets is associated with skin barrier dysfunction in children with atopic dermatitis. Clinical and Experimental Allergy. 2021; 51(5):666-673.
CYP2D6 Phenotype Influences Aripiprazole Tolerability in Pediatric Patients with Mood Disorders. Journal of Child and Adolescent Psychopharmacology. 2021; 31(1):56-62.
Very early onset eosinophilic esophagitis is common, responds to standard therapy, and demonstrates enrichment for CAPN14 genetic variants. Journal of Allergy and Clinical Immunology. 2021; 147(1):244-254.e6.
From the Blog
Yes, Genetic Disease Risk Information Can Be Shared Responsibly
Lisa J. Martin, PhD, Leah C. Kottyan, PhD3/23/2026
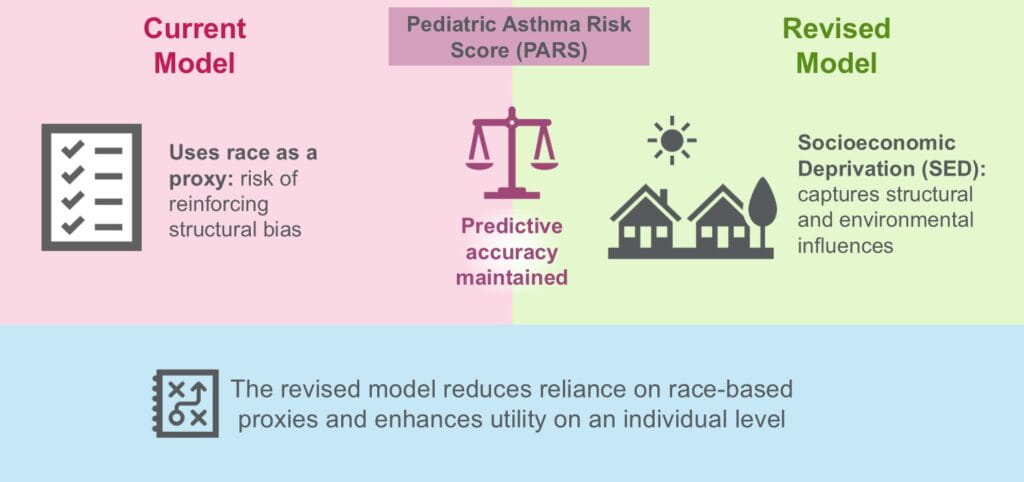
A ‘Must Read’—An Update on Asthma Prediction
Lisa J. Martin, PhD, Jocelyn M. Biagini, PhD ...1/21/2026
Parents Can Reliably Report Their Child’s EoE Symptoms Over Time
Lisa J. Martin, PhD12/19/2024