About
Biography
As a molecular epidemiologist, I am engaged in researching pediatric allergic diseases. My special interests include asthma and food allergy, allergic disease progression, and the genomic and environmental contributions to allergic diseases in children. I was drawn to these research areas after growing up with seasonal allergies and asthma, and by my love of pediatric research.
In the Biagini Lab, our long-term goals are to improve the health of children by conducting research that advances the field of asthma and allergic disease research by investigating the interplay of genetics with environmental exposures. We are also studying the mechanisms underlying the progression of disease from food allergy to asthma and the biology underlying severe asthma in children.
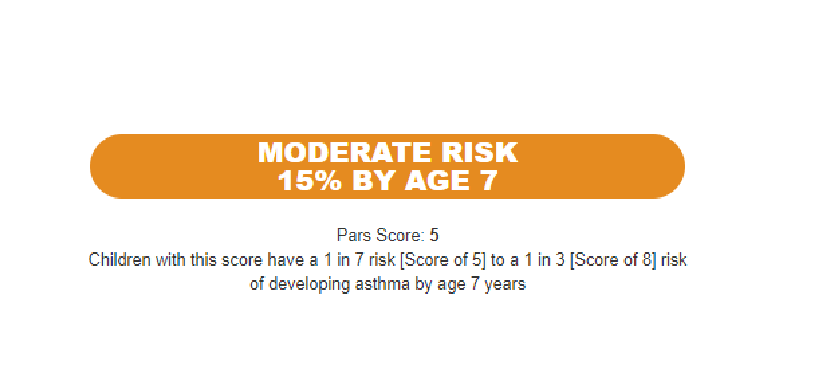
We recently published the Pediatric Asthma Risk Score (PARS), which is the first continuous risk score for asthma in children. It relies on factors that are routinely collected in the assessment of a child being evaluated for allergy, asthma or both. It is superior to the traditional Asthma Predictive Index (API) with an 11% increase in sensitivity due to improved prediction in children with mild-to-moderate asthma risk.
I am an elected member to the American Academy of Asthma, Allergy and Immunology and a member of the editorial board for the Journal of Asthma. I began my career in 2001 and joined the Cincinnati Children’s team in 2008.
BS: Xavier University, Cincinnati, OH, 2001
MS: University of Cincinnati, Cincinnati, OH, 2004
PhD: University of Cincinnati, Cincinnati, OH, 2008
Postdoctoral Fellowship: Division of Asthma Research, Cincinnati Children's Hospital Medical Center, Cincinnati, OH, 2011
Research Areas
Publications
A Pediatric Asthma Risk Score to better predict asthma development in young children. Journal of Allergy and Clinical Immunology. 2019; 143(5):1803-1810.e2.
Redlining, Community Wealth, and Air Pollution: A Tale of Three Cities-Boston, Nashville, and Detroit. Journal of Urban Health. 2026.
Longitudinal enrichment of eosinophilic esophagitis in children with AD: The MPAACH cohort. Journal of Allergy and Clinical Immunology. 2026.
B-Cell Repertoire of Children With Atopic Dermatitis Exhibits Altered IgE Maturation Associated With Allergic Food Sensitization. Allergy. European Journal of Allergy and Clinical Immunology. 2026; 81(2):513-523.
Genome-wide cross-trait analysis of heterogeneous outcomes in early life atopic dermatitis. Pediatric Allergy and Immunology. 2025; 36(12):e70253.
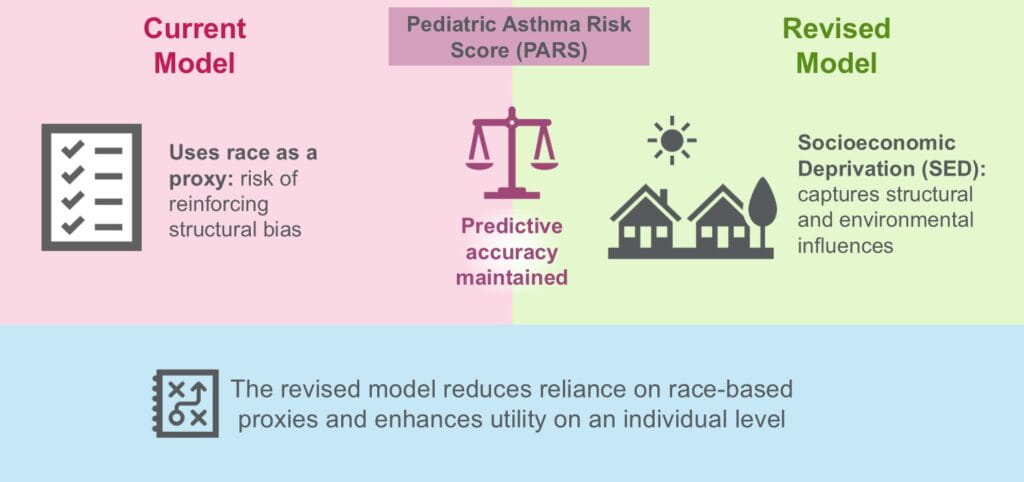
Socioeconomic deprivation performs equally to race in the Pediatric Asthma Risk Score. Pediatric Allergy and Immunology. 2025; 36(12):e70263.
Objective sensitization algorithm validates parental report of food allergy in children. Journal of Allergy and Clinical Immunology: Global. 2025; 4(4):100512.
Risk Factors and Mechanisms Leading to Preschool Recurrent Wheeze and Asthma. Journal of Allergy and Clinical Immunology: In Practice. 2025; 13(10):2537-2551.
Early-life wheeze trajectories are associated with distinct asthma transcriptomes later in life. Journal of Allergy and Clinical Immunology. 2025; 156(3):640-650.
Phenotypes of Atopic Dermatitis and Development of Allergic Diseases. JAMA Network Open. 2025; 8(6):e2515094.
From the Blog
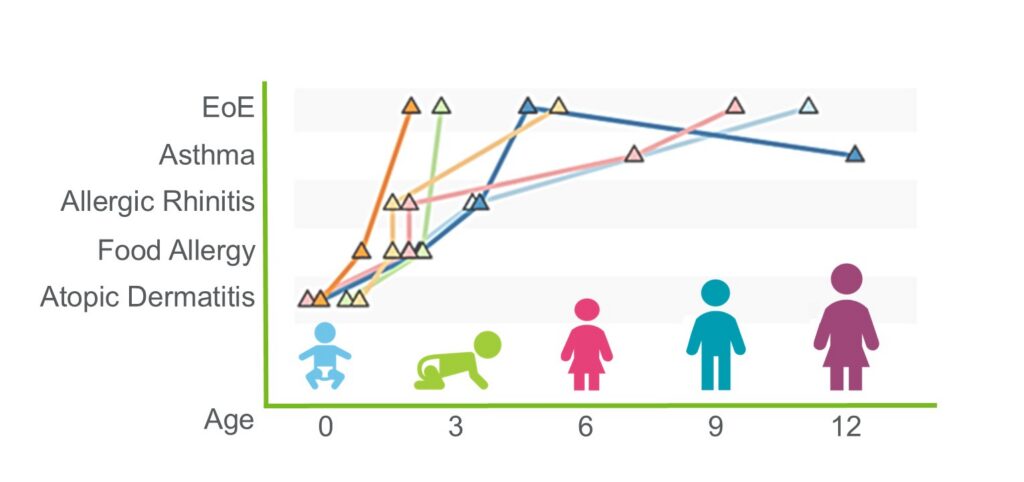
EoE: The Fifth Member of the Atopic March
Jocelyn M. Biagini, PhD, Marc E. Rothenberg, MD, PhD ...3/31/2026
A ‘Must Read’—An Update on Asthma Prediction
Jocelyn M. Biagini, PhD, Lisa J. Martin, PhD ...1/21/2026
Omitting Race from Lung Function Equations Increases Detection of Asthma in Black Children
Jocelyn M. Biagini, PhD, Gurjit Khurana Hershey, MD, PhD2/28/2025
Asthma Risk Score Tool Performs Well Across Diverse Populations
Jocelyn M. Biagini, PhD, Gurjit Khurana Hershey, MD, PhD8/11/2023
Compromised Skin Outweighs Inheritance as Allergy Risk
Jocelyn M. Biagini, PhD2/18/2020