About
Biography
I am interested in the physical and functional organization of the cell nucleus. My current research program focuses on two research areas: (1) fundamental mechanisms regulating gene expression and chromosome organization and (2) human diseases caused by dysfunctions of the nuclear envelope such as cardiomyopathies and premature aging disorder progeria.
Our lab investigates mechanisms of gene transcription during the cell cycle, cell quiescence, cell differentiation and disease processes. We investigate the relationship between gene transcription and transcription factor binding, chromatin accessibility, large-scale chromosome organization, and subnuclear localization of genomic regions to the nuclear pore and the nuclear lamina. We employ powerful genomics tools combined with bioinformatics, cell engineering and microscopy. We also develop new technologies to investigate mechanisms of gene transcription. Our goal is to obtain a quantitative and system-level understanding of gene expression and chromosome organization within the nucleus.
Our lab extensively studies the pathogenesis of laminopathies, a spectrum of degenerative diseases caused by mutations in the genes encoding nuclear lamins. Nuclear lamins are components of the nuclear envelope. Laminopathies affect both children and adults and include dilated cardiomyopathy, Emery-Dreifuss muscular dystrophy, familial partial lipodystrophy and the premature-aging disorder, Hutchinson-Gilford progeria. To identify the pathogenic components of laminopathies, we examine various biological functions of nuclear lamins, including their roles in chromosome organization, gene regulation and physical protection of the cell nucleus. We currently focus on the etiology of dilated cardiomyopathies and progeria. In this research project, we use cardiac in vivo pathophysiological analyses, cells derived from laminopathy patients, induced pluripotent stem cells, mouse models of laminopathies and functional genomics techniques. The long-term goal of this project is to identify molecular pathways that can be targeted to halt or significantly delay the progression of laminopathies.
My research discoveries include: phosphorylated Lamin A/C binding to gene enhancers and its disruption in progeria (PMID: 32208162); nuclear pore interactions with RNA Polymerase III-transcribed genes (PMID: 24011592); and nuclear lamina-associated domains in the C. elegans genome (PMID: 21176223). In addition, I produced numerous public genomic data sets for chromatin protein binding sites and histone modifications in C. elegans as a member of the modENCODE consortium (PMID: 21177976; PMID: 25164756).
After receiving my PhD at the University of Tokyo, I worked as a postdoctoral researcher at the Department of Biology at the University of North Carolina at Chapel Hill, and then as a research scholar at Lewis-Sigler Institute for Integrative Genomics at Princeton University. Before joining Cincinnati Children’s in 2021, I worked as a Pathway-to-Independence track instructor at the Department of Pediatrics at the University of Chicago. I received the Tanabata Fellowship from RIKEN Genomic Sciences Center (2010) and the Best Basic Research Award for Junior Faculty in the Department of Pediatrics at the University of Chicago (2017, 2018, 2019). I have been a researcher for over two decades.
BS: Tamagawa University, Tokyo, Japan, 2002
PhD: The University of Tokyo, Tokyo, Japan, 2007
Postdoctoral: The University of North Carolina at Chapel Hill, Chapel Hill, NC, 2012
Interests
Cardiovascular diseases; laminopathies; progeria; chromosome organization; transcriptional regulation; nuclear envelope
Research Areas
Publications
Starvation activates ECM-remodeling gene transcription and putative enhancers in fibroblasts despite inducing quiescence. Cell reports. 2025; 44(7):115896.
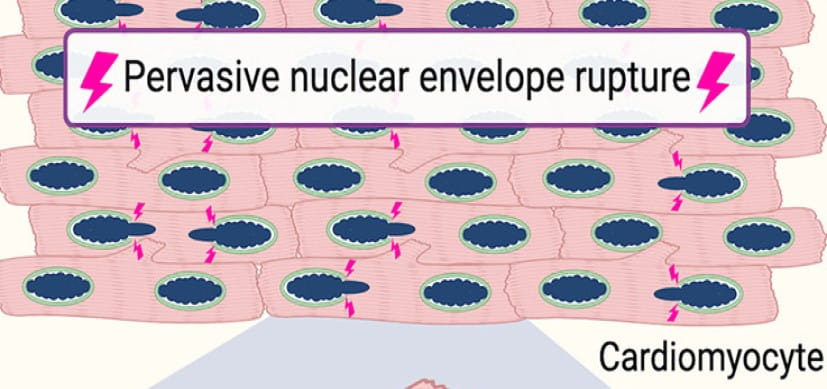
Pervasive nuclear envelope ruptures precede ECM signaling and disease onset without activating cGAS-STING in Lamin-cardiomyopathy mice. Cell reports. 2024; 43(6):114284.
ETV2 primes hematoendothelial gene enhancers prior to hematoendothelial fate commitment. Cell reports. 2023; 42(6):112665.
Hedgehog signaling activates a mammalian heterochronic gene regulatory network controlling differentiation timing across lineages. Developmental Cell. 2022; 57(18):2181-2203.e9.
Nuclear lamin phosphorylation: an emerging role in gene regulation and pathogenesis of laminopathies. Nucleus. 2020; 11(1):299-314.
Phosphorylated Lamin A/C in the Nuclear Interior Binds Active Enhancers Associated with Abnormal Transcription in Progeria. Developmental Cell. 2020; 52(6):699-713.e11.
TRA-1 ChIP-seq reveals regulators of sexual differentiation and multilevel feedback in nematode sex determination. Proceedings of the National Academy of Sciences of the United States of America. 2013; 110(40):16033-16038.
Integral nuclear pore proteins bind to Pol III-transcribed genes and are required for Pol III transcript processing in C. elegans. Molecular Cell. 2013; 51(6):840-849.
Broad chromosomal domains of histone modification patterns in C. elegans. Genome Research. 2011; 21(2):227-236.
From the Blog
Study Challenges Current Thinking About Mechanisms of Dilated Cardiomyopathy
Kohta Ikegami, PhD6/11/2024