Single Cell Applications in Lymphangioleiomyomatosis (LAM)
Lymphangioleiomyomatosis (LAM) is a progressive and often fatal lung disease affecting primarily women of childbearing age. LAM is associated with mutations in the tuberous sclerosis genes, TSC1 or TSC2, resulting in activation of the mechanistic target of sirolimus complex 1 (mTORC1). One of the biggest challenges in the development of optimal therapeutic strategies for LAM is the lack of understanding of the origin of the LAM cells and the mechanisms contributing to LAM cell proliferation, migration and metastasis. Our goal is to address the fundamental questions of 1) what are LAM “cells”; 2) where does it come from; and 3) how are the LAM “cells” regulated using single cell approaches.
- Identification and characterization of the lymphangioleiomyomatosis (LAM) cell and its origin using integrative analysis of single cell RNA-seq and single cell ATAC-seq
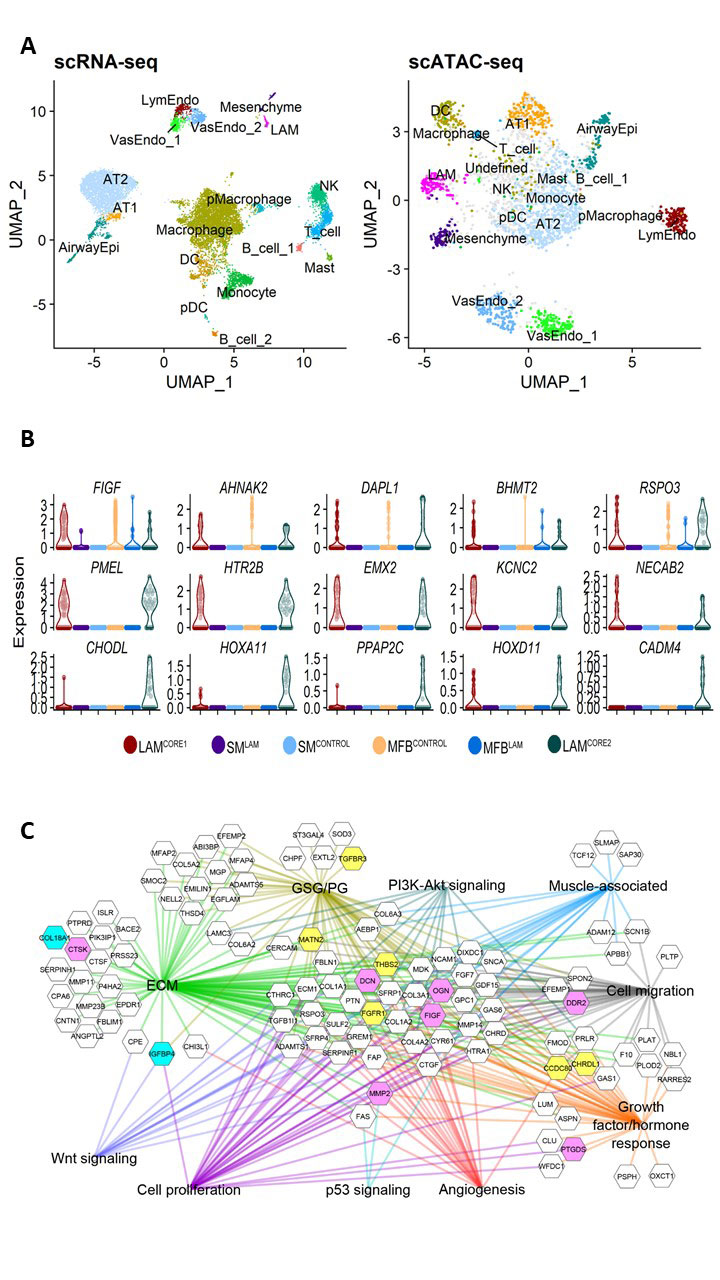
We performed single cell and single nuclei RNA-seq (scRNA-seq) on LAM lung and uterus, and compared with normal lung and normal uterus. Our single cell transcriptomic analysis identified a unique population of cells termed LAMCORE in LAM lung and uterus, demonstrated their transcriptomic similarity, and predicted novel signature genes and altered pathways in LAM (Guo et al., bioRxiv 784199). Single-cell ATAC-seq (scATAC-seq) was recently performed to assess genome-wide chromatin accessibility profiling of LAM and normal lungs. Integrative scRNA and scATAC analysis is ongoing, the study will correlate chromatin accessibility states and gene expression of LAM cells, provide further understanding of the genomic and epigenetic regulation of LAM pathogenesis.
- The homeobox gene network and its role in pulmonary lymphangioleiomyomatosis (LAM) pathogenesis
While the role of female steroid hormones in the pathogenesis of LAM is well established, lack of detailed mechanistic understanding limits the development of new therapies for LAM patients. LAM single cell and single nuclei RNA sequencing (scRNA-seq & snRNA-seq) analysis identified a unique population of pulmonary LAMCORE cells expressing uterine-specific HOXs that are rarely detectable in normal lung. The role of HOX family of transcription factors on the pulmonary LAM pathogenesis is unknown. How LAMCORE cell-specific HOX gene network mediates genomic and epigenetic regulation of on LAM cell migration, metastasis, and trans-differentiation are unclear. We will employ scRNA-seq and scATAC-seq to interrogate genomic and epigenetic effects simultaneously to reveal estrogen/progesterone-driven HOXs signaling in LAM; to identify the HOXs centered gene network and its regulatory roles in LAM progression and to test the effect of the HOX antagonist, HXR9, singly or in combination with Sirolimus, on estrogen and/or progesterone-promoted LAM metastasis and lung remodeling in vivo in preclinical models to address questions of direct clinical relevance.
Grant Support
LAM Foundation Established Investigator Award (Perl, Xu)
1R01HL153045-01 (Xu and Yu)