Brian Gebelein, PhD
- Director, MDB graduate program
- Professor, UC Department of Pediatrics
About
Biography
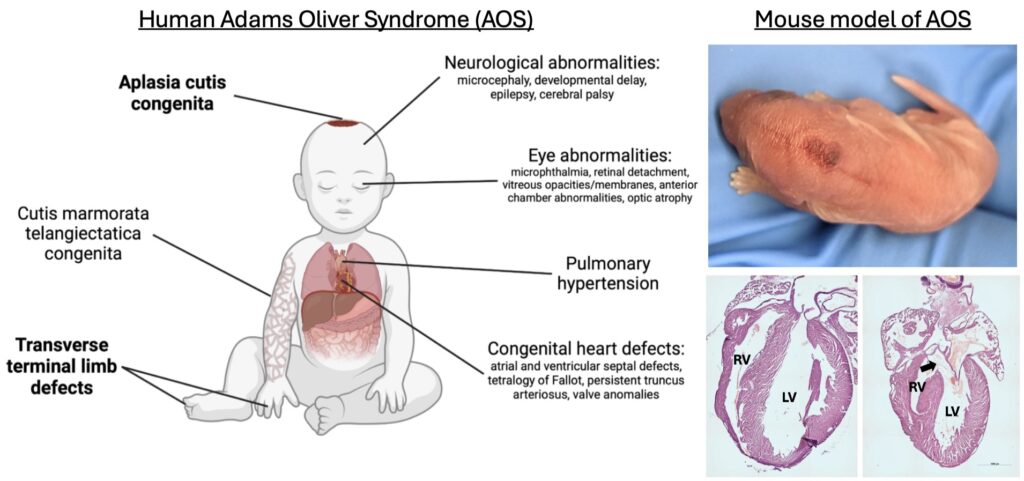
My research areas include developmental biology, gene expression regulation, neural cell-fate specification, mechanisms patterning the early embryo and the mechanisms underlying Notch signaling, with a specific emphasis on modeling the disease alleles of Adams-Oliver syndrome.
The goal of my laboratory is to understand the molecular mechanisms underlying cell-specific transcription. I have always been fascinated with how cells within our body can use the same genome to generate the vast array of cell types needed to form functional organs and tissues.
Some of my groundbreaking discoveries include:
- How Hox transcription factors are integrated with genes regulating segmentation to pattern the early embryo
- How Hox transcription factors are integrated with zinc finger proteins to pattern the nervous system and regulate hematopoiesis
I received the Basil O'Connor Award from the March of Dimes in 2006. I have been a researcher for more than 26 years and began my work at Cincinnati Children's in 2004. My research has been published in respected journals, such as eLife, Nature, Science, Developmental Cell, Genes and Development, Development, Developmental Biology and PLoS Genetics.
BS: University of Wisconsin, Milwaukee, WI, 1994
PhD: Mayo Graduate School, Rochester, MN, 2000
Fellowship: Molecular mechanisms of Hox specificity in Drosophila melanogaster, Columbia University
Interests
Developmental genetics; Transcriptional regulation; Modeling disease alleles in mice, Drosophila, and human stem cells.
Publications
An inducible system to study the regulatory functions of GSX2 in human lateral ganglionic eminence-like progenitors. Developmental Biology. 2026; 532:35-51.
A dual role for GLI3 signaling in neural crest development. Development. 2026; 153(1).
Defective Notch1 signaling in endothelial cells drives pathogenesis in a mouse model of Adams-Oliver syndrome. Journal of Clinical Investigation. 2025; 135(23).
The ALX4 dimer structure provides insight into how disease alleles impact function. Nature Communications. 2025; 16(1):4800.
Modelling a pathological GSX2 variant that selectively alters DNA binding reveals hypomorphic mouse brain defects. Disease Models & Mechanisms. 2025; 18(2).
Cooperative Gsx2-DNA binding requires DNA bending and a novel Gsx2 homeodomain interface. Nucleic Acids Research (NAR). 2024; 52(13):7987-8002.
Homeodomain complex formation and biomolecular condensates in Hox gene regulation. Seminars in Cell and Developmental Biology. 2024; 152-153:93-100.
Prediction of cooperative homeodomain DNA binding sites from high-throughput-SELEX data. Nucleic Acids Research (NAR). 2023; 51(12):6055-6072.
A Drosophila Su(H) model of Adams-Oliver Syndrome reveals cofactor titration as a mechanism underlying developmental defects. PLoS Genetics. 2022; 18(8):e1010335.
Olig2 defines a subset of neural stem cells that produce specific olfactory bulb interneuron subtypes in the subventricular zone of adult mice. Development. 2022; 149(5).
From the Blog
Genetic Research Pinpoints a Lethal Risk Linked to Adams-Oliver Syndrome
Brian Gebelein, PhD12/1/2025