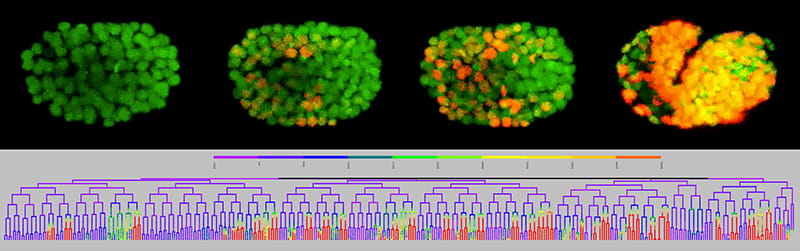
Model System
We utilize an innovative live-imaging approach that lets us measure the expression of a gene or other reporter with single cell resolution in all cells throughout development. We study the development of the tiny, but highly conserved, nematode, C. elegans, an excellent genetic model system. When combined, our approach and model system have some powerful scientific advantages.
What is C. elegans?
Caenorhabditis elegans is a free-living (non-parasitic) worm approximately 1 mm long. They are visible to the naked eye but we use microscopes to visualize them better—both the embryo and adult are transparent. In the wild, they eat the bacteria that grow in rotting fruit and vegetation; in the lab, we grow them on agar plates with their food source, so they are very inexpensive to maintain. C. elegans are self-fertilizing hermaphrodites that complete the life cycle from fertilized egg to fertile adult in only three days, and males occur occasionally in the population which makes genetic crosses easy and fast. We can even freeze strains to save for future use.
C. elegans was the first animal to have its complete genome sequenced in 1999. It has just over 20,000 genes, similar to the number of genes in the human genome and many of these genes are conserved by evolution, particularly those involved in development. The adult C. elegans has only 959 somatic cells, which include neurons, muscle, skin, and intestinal cells with similar functions and gene expression to those found in humans.
What is the advantage of C. elegans in developmental biology?
For developmental biologists, the biggest advantage of C. elegans is its “invariant lineage”. This means that every embryo develops with the exact same pattern of cell divisions and cell migrations and hatches with the exact same complement of 558 cells. Larger, more complex embryos like vertebrates have millions more cells, so their development is more variable; the small size of the worm embryo both enables and requires precision. Because of the invariant lineage, we can predict the future for a given cell: we know what fate it will adopt before it starts to express any markers of its fate.