Current Projects
 Myeloid cells in inflammation and obesity
Myeloid cells in inflammation and obesity
The biological systems of inflammation and metabolism are closely intertwined, and tissue macrophages are key immune cells linking these systems. In addition to their roles as sentinels of infection, macrophages also contribute to obesity-associated diseases.
For example, certain diets can drive tissue accumulation of inflammatory lipids and augment intestinal permeability. These inflammatory lipid species and products of the leaky gut may result in local and systemic inflammation.
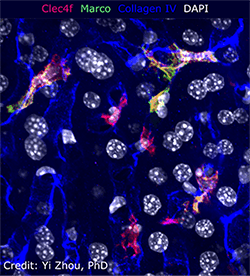
One documented consequence is the development of insulin resistance and increased obesity, driven by adipose tissue – recruited macrophages. Additionally, hepatic macrophages lining the liver sinusoids become more susceptible to death and inflammatory dysregulation. These hepatic macrophages function to clear microbes and microbial products draining from the intestine, and absence and dysregulation of these hepatic macrophages may exacerbate systemic inflammation.
One area of interest in the Troutman Lab is to define molecular mechanisms controlling tissue macrophage behaviors resulting from inflammation and obesity.
Genetic variation and control of macrophage behaviors
Normal tissue behaviors depend on a network of cell populations functioning together to support organ function. Disease may result if the system becomes unbalanced in one or more constituent cell types. Some diseases have known lesions in a single causal gene. However, most diseases result from the polygenic background of a susceptible individual interacting with a range of environmental factors. Thus, the susceptibility of a tissue to develop disease may be governed by interactions of the genetic background with one or more environmental factors. Notable examples include Alzheimer disease, cardiovascular disease, insulin resistance, steatohepatitis and cirrhosis and atopic diseases. In these cases, certain populations are resistant to disease progression in the context of similar environmental triggers.
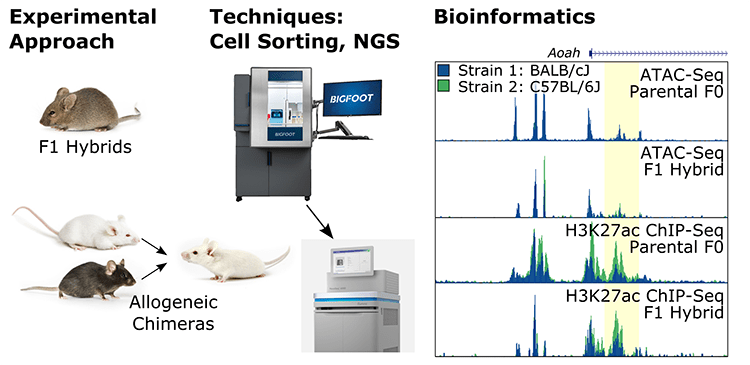
Genome-wide association studies provide the identities of susceptibility loci for many complex diseases. These susceptibility loci often lie in noncoding regions of the genome, which suggest that they function by regulating gene expression. However, different cell lineages can select divergent sets of noncoding regulatory elements controlling cell fate and function. Therefore, improving mechanistic understanding of the genetic and environmental factors interacting to promote disease may require knowledge of the cell types actively using the mutated DNA elements. The Troutman Lab studies these interactions using purified cells from inbred mouse strains. Gene regulatory differences between the model genomes are assessed using experimental conditions that fix the contribution of environmental effects. This enables defining cell autonomous and non–cell autonomous roles for naturally occurring genetic variation in controlling macrophage behaviors. Although the locations of gene regulatory elements have diverged between humans and mice, the mechanisms controlling gene regulation and the functional properties of macrophages remain highly conserved between the two species. Thus, the prevalence of mutations in mouse macrophages affecting the binding of a certain transcription factor may cause similar outcomes in humans.